In silico binding studies of NodD proteins of Bradyrhizobium yuanmingense and Rhizobium tropici with flavonoids
DOI:
https://doi.org/10.5530/ctbp.2026.2.20Keywords:
NodD protein, Bradhyrhizobium yuanmingense, Rhizobium tropici, Flavonoids, Molecular modeling, Docking, MD simulationsAbstract
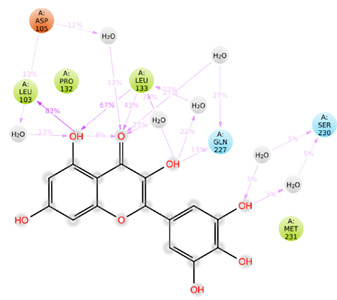
Prokaryotic bacteria, such as Rhizobium engage in endosymbiosis with legume plants’ roots. In this mutualistic relationship, bacteria obtain carbon from the plant, while the plant benefits from nitrogen-fixation. The formation of root nodules in leguminous plants is induced by flavonoids exuded by the plant roots, acting as chemo-attractants for symbiotic rhizobia. The interaction between these flavonoids and NodD proteins is crucial for the induction of nodulation genes, influencing host range specificity. The initial signal exchanged during nodule development involves the interaction of legume root-exuded flavonoids with rhizobial NodD proteins, representing an early checkpoint in the establishment of symbiosis. In this current work, we focused on studying the molecular level interactions of flavonoids, common inducers of other rhizobial NodD protein with F1JZX9 protein (Nod D1) of Bradyrhizobium yuanmingense along with Q02876 protein (Nod D1) and P32008 protein (Nod D2) of Rhizobium tropici by In-silico studies. Docking studies, sequence alignment analysis, and molecular dynamics simulation studies were all applied in the in-silico investigations of molecular level interactions of flavonoids with common inducers of other rhizobial NodD protein of Bradyrhizobium yuanmingense and Rhizobium tropici. From our In Silico analysis, it was revealed that Flavonoids Myricetin and Daidzen have the highest potential to induce the nodulation process via binding to Nod D1 and Nod D2 proteins respectively. This molecular level understanding of how these flavonoids are selectively attracted towards NodD proteins can help in prioritizing the most binding efficient flavonoid towards attaining sustainable agriculture.