Genome-wide Study and Expression Profiles of a Promising Negative Regulator of Abiotic Stress Gene Family OsDHSRP in Oryza sativa
DOI:
https://doi.org/10.5530/ctbp.2026.2.15Keywords:
Abiotic stress, Climate resilient crops, E3-ubiquitin ligase, OsDHSRP, Oryza sativaAbstract
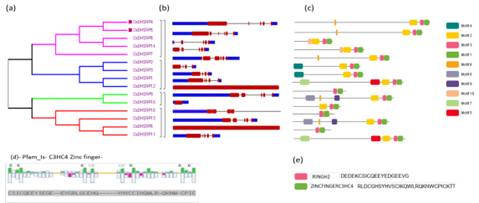
Rice is the global staple food, but the crop has been gradually impeded by many environmental constraints such as drought, floods, salinity, heat, and cold. Generally, plants adapt to environmental cues by altering physiological conditions primarily by modulating the expression of several stress-responsive genes. OsDHSRP1 is an E3-ubiquitin ligase whose expression is highly stimulated by salinity, heat, and drought conditions, and it acts as a negative modulator by boosting reactive oxygen species (ROS) production, thereby causes damages to vital cellular organelles. In the present study, we are reporting genome-wide prediction, structural, evolutionary characterization, and expression analysis of OsDHSRP gene family of Oryza sativa under diverse abiotic stress conditions. A total of 15 OsDHSRP genes were discovered in Oryza sativa genome, which contains a C3HC4 zinc finger conserved domain. The elucidation of intron/exon and motif patterns provides structural aspects of these genes. Cis-regulatory analysis and transcription factor prediction studies revealed their roles and interactions with genes involved in multiple abiotic variables. Expression analysis of OsDHSRP genes by RT-qPCR revealed that OsDHSRP1 is associated with cold, drought and salt stress conditions, suggesting the role of OsDHSRP1 during diverse abiotic stresses. This study provides further insights into the regulation of expression of OsDHSRP genes for developing climate resilient crops.